Nutrigenomics and Public Health
Nutrigenomics is a relatively new and very fast-moving field of research which combines molecular biology, genetics, and nutrition. It focuses on the role of nutritional status or specific nutrients in the regulation of gene expression (Afman and Muller, 2006; Komduur et al., 2009; Mead, 2007). While one of its effects may be the development of personal diets based on an individual’s genetic code and focused on optimizing the expression of certain genes therein, other health-related outcomes show more promise at this juncture and have greater implications for public health promotion (Ordovas and Corella, 2004; Muller and Kersten, 2003). Of significant interest is the potential for greater knowledge of how diet may influence disease state at the gene or molecular levels. Many threats to public health, including cancer, diabetes, obesity and other chronic diseases, are influenced to a certain extent by genetic factors. The prevalence of cardiovascular disease, diabetes, and other related conditions in the United States is shown in Figure 1.
 Traditionally, many recommendations for diet intake have been population based due to findings from epidemiology studies. For instance, recommendations to lower fat intake for a population are due to a link between fat intake and heart disease. However, not all individuals who consume a high-fat diet will develop heart disease. If we know who has a genetic susceptibility to a type of heart disease associated with elevated blood cholesterol, then perhaps diets can be tailored to those individuals. With the completion of the human genome project and continuous identification of the role of human genes, more information on who may be more likely to develop certain diseases and whether diet can prevent the expression of these diseases is an area of fundamental research.
Traditionally, many recommendations for diet intake have been population based due to findings from epidemiology studies. For instance, recommendations to lower fat intake for a population are due to a link between fat intake and heart disease. However, not all individuals who consume a high-fat diet will develop heart disease. If we know who has a genetic susceptibility to a type of heart disease associated with elevated blood cholesterol, then perhaps diets can be tailored to those individuals. With the completion of the human genome project and continuous identification of the role of human genes, more information on who may be more likely to develop certain diseases and whether diet can prevent the expression of these diseases is an area of fundamental research.
The impact of nutrigenomics is not limited to public health. The food science and culinary industries may benefit from the information revealed by nutrigenomic research, and may also take advantage of opportunities to promote public health in a more active way. Understanding the impact of processing on food composition, added to new information regarding nutrients’ effects on genetics and disease processes, will afford new chances to improve public health through everyday manufacturing and food-processing practices. One such opportunity is the further development of functional foods with higher nutritional and sensory quality (Ferguson, 2009).
This article will focus on the potential public health impact of information gleaned from nutrigenomic research. We will explore how genes affect disease, and the ways in which nutrients affect genes. We will then discuss the potential impact of nutrient intake on disease state in light of current food consumption trends. Finally, we will outline how the food and culinary industries may take advantage of this information, and opportunities for exerting a positive influence on public health.
Genetic Regulation of Disease State
It has long been recognized that a family history of various chronic diseases increases an individual’s risk for developing such diseases. In general, the closer the genetic relationship between a patient and an affected family member, the greater the risk that the patient might develop the disease in question. The best explanation for this phenomenon is that many chronic diseases are regulated not only by lifestyle factors and environmental influences, but also by some genetic component.
Since the advent of sensitive molecular biology procedures, many genes partially responsible for the development of chronic disease have been suggested, and some have been confirmed. Among these, perhaps the most well known are the oncogenes. Oncogenes are genes which code for proteins that control cell death, or apoptosis; or which control cell division and proliferation (Croce, 2008). Rapid cell proliferation or inhibition of apoptosis may cause a cell to become cancerous, and begin the growth of lesions which may, in turn, become tumors. Which mechanism causes the onset of cancer is dependent on the specific oncogene and tissue type involved (Land et al., 1983).
--- PAGE BREAK ---
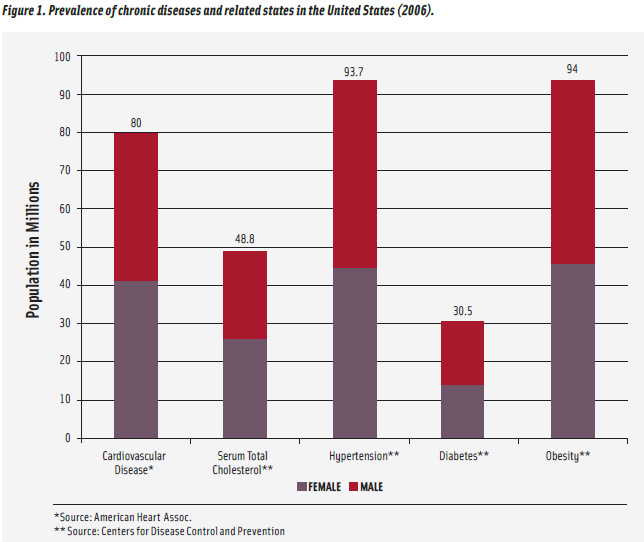 Another chronic disease with vast implications for public health is cardiovascular disease. Common biochemical markers for elevated cardiovascular risk include nonspecific indices of systemic inflammation, and their elevation is associated with cardiovascular risk, although they may be elevated due to more acute conditions such as infection. It has been demonstrated that the circulating levels of these markers may be influenced, at least in part, by various genetic factors (Pankow et al., 2001). Other genetic factors have also shown an effect in regulating the overall energy balance of an individual, contributing to the degree of one’s obesity or leanness (Barsh et al., 2000). These findings may influence how cardiovascular disease risk is diagnosed.
Another chronic disease with vast implications for public health is cardiovascular disease. Common biochemical markers for elevated cardiovascular risk include nonspecific indices of systemic inflammation, and their elevation is associated with cardiovascular risk, although they may be elevated due to more acute conditions such as infection. It has been demonstrated that the circulating levels of these markers may be influenced, at least in part, by various genetic factors (Pankow et al., 2001). Other genetic factors have also shown an effect in regulating the overall energy balance of an individual, contributing to the degree of one’s obesity or leanness (Barsh et al., 2000). These findings may influence how cardiovascular disease risk is diagnosed.
Hyperinsulinemia and diabetes are examples of chronic conditions plaguing the general public which have their origins, or partial origins, in genetics. Defective genetic regulation of several proteins which respond to the presence of insulin may be implicated in the etiology of type 2 diabetes. Ducluzeau et al. (2001) found decreased expression of key proteins and their mRNA in type 2 diabetic patients, and observed a lack of response to the presence of insulin by up-regulation of key response proteins. This lack of response is linked to defective expression of the genes for these proteins in response to insulin. Type 1 diabetes is reliant on genetic components to a greater degree than type 2 diabetes because the genetic regulation of type 1 diabetes is polygenic, or accomplished by several genes.
Nutrient Regulation of Gene Expression
In some cases, intake or levels of specific nutrients have been shown to directly impact gene expression. Often, this occurs by means of transcription factors. Transcription factors are biochemical entities which bind to the DNA and either promote or inhibit transcription of genes. Several nutrients are known to bind to transcription factors and regulate gene expression in this manner (Muller and Kersten, 2003).
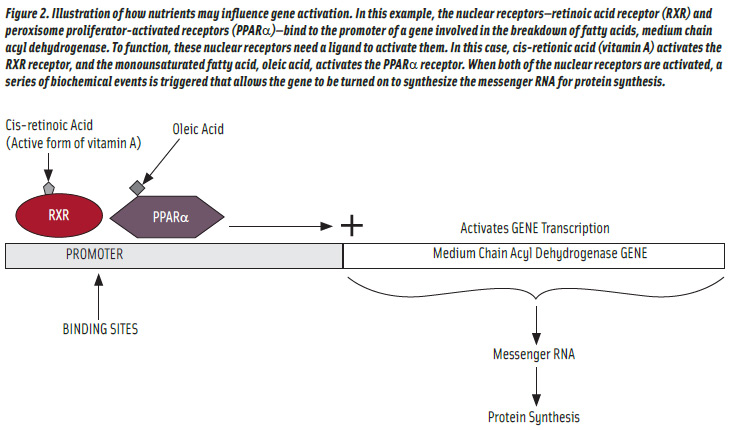
The regulation of gene transcription by fatty acids is perhaps the best understood of the nutrient gene-regulation pathways. A major pathway by which fatty acids modulate gene expression is the peroxisome proliferator-activated receptors (PPARs), which activate gene transcription when fatty acids bind to them (Bouwens et al., 2007). One such receptor has been shown to activate genes responsible for utilization of fatty acids for energy (Kresten et al., 1999). Circulating fatty acid concentrations are increased during fasting. These fasting-induced amplified fatty acid concentrations increase the expression of several genes involved in β-oxidation of fatty acids (Bouwens et al., 2007). One of the isoforms of PPAR is PPARα, which is a nuclear receptor that binds to the promoters of genes whose products favor fatty acid breakdown (Madrazo and Kelly, 2008). However, these receptors need activators to realize their full potential. Fatty acids can act as ligands to PPARα, such as oleic acid, to enhance activity (Michaud and Renier, 2009). Furthermore, the PPARs do not act by themselves but require another partner, the retinoic acid receptors with cis-retinoic acid as a ligand. Thus, fatty acid catabolism is somewhat dependent upon cis-retinoic acid or any other nutrient or drug that enhances the activity of the nuclear receptors (Figure 2). Also, PPAR activity activation may lead to weight loss (Verreth et al., 2004). However, much has been done in developing drugs to act as ligands to both PPARα and PPARγ; perhaps the same approach may be taken by the food industry (Robinson and Grieve, 2009).
Dietary carbohydrate intake has also been implicated in the regulation of gene expression. Glucose consumption is a crucial determinant of the rate of transcription in the liver of genes for several glycolytic enzymes and enzymes for fatty acid synthesis, as well as lipoproteins and insulin (Clarke and Abraham, 1992). However, this effect has also been observed in other tissues such as the pancreas (German et al., 1990).
--- PAGE BREAK ---
Vitamins and minerals may also have transcriptional regulation properties. Retinoic acid, a form of vitamin A, forms part of transcription factors which may bind to the promoter region of genes involved in embryonic development and cell differentiation (DeLuca, 1991). Divalent minerals, such as iron, copper, and zinc, alter the transcription rates of genes for proteins such as metallothionein, a crucial transport and storage protein for these minerals (Bremener and Beattie, 1990). Probably the most notable nutrient gene interaction deals with vitamin D and the vitamin D nuclear receptor. Here, 1, 25 dihydroxycholecalciferol binds with the nuclear receptor, and in the nucleus this complex binds to promoters of genes that encode for many proteins. In essence, the role of vitamin D appears to favor pathways involved with cell differentiation and inhibits pathways avoring cell proliferation. Hence, this may have an impact on cancer as cell proliferation versus differentiation must be kept in balance (Samuel and Sitrin, 2008).
Folate metabolism may have variations among populations. When there is a mutation in one nucleotide base pair that alters the function of the protein that occurs in more than 1% of the population, this is termed a single nucleotide polymorphism or SNP. Folate synthesis in some populations may be limited due to SNP and thus more dietary folate may be required (Martínez de Villarreal, 2001). A lack of folate during pregnancy can lead to spina bifida birth defects. A Mexican study suggested an increased risk of colorectal cancer with a particular type of polymorphism (Gallegos-Arreola et al., 2009). Diets customized to provide extra folate to those with these SNP profile could be developed. Overall, food may impact the cell by direct interaction with DNA (genes), RNA, protein, or other metabolites. Knowing these roles of nutrients and how each of these cell components may cause diseases will allow food to be tailored to an individual’s genetic make-up.
Nutrigenomics and Food Processing
Having discussed the ability of various nutrients to impact gene expression and the ability of genes to influence disease progression, examination of the impacts of processing techniques on the nutrient content or quality of several foods is appropriate. It may be that food processing or preparation practices can change levels of nutrients which will impact genes regulating disease processes. If this is the case, consideration of current industry practices may be in order. In general, most food preservation processes lead to diminished nutritional quality for food products, of course, the challenge being to balance food safety issues with nutritional aspects.
Traditional methods such as canning and freezing have been shown to have detrimental effects on nutritional quality of food products (Schroeder, 1971; Hale and Brown, 1983; Aubourg, 1997; Slavin et al., 2001). Minimal processing techniques are becoming more popular, especially with traditional and healthy food like fruits and vegetables (Allende et al., 2006). Other non-conventional techniques such as osmotic dehydration, membrane processing, ultra high hydrostatic pressure, and high voltage pulsed electric fields to name a few are being explored more and more for food processing applications (Ortega-Rivas, 2007) to better preserve the nutritional quality of processed foods.
--- PAGE BREAK ---
Implications for the Food and Culinary Industries
How can nutrigenomic-related research and findings be applied by the food and culinary industries to promote health? A consumer acceptance study about nutrigenomics by Ronteltap et al. (2009) showed that people are open to this concept, provided that there is more communication between the experts and consumers, and also more information should be made available to them. Although genetically sensitive foods are different from genetically modified foods, consumer skepticism cannot be ruled out if the information about nutrigenomics is not conveyed properly to consumers (Archibald, 2004). According to Archibald (2004), a genetic counselor or a physician could help an individual understand his/her genetic makeup, and then the nutritional professional could tailor diets accordingly (Figure 3).
Let us suppose for example that since oleic acid enhances PPARα activity and therefore fatty acid oxidation, should food and culinary entities choose food high in oleic acid (i.e., peanut or olive oils) in some of their meals or menus? Dysfunctional PPARα activity has been implicated in diabetes and thus it is not inconceivable that providing a diet to enhance PPARα activity may prove beneficial. Indeed, oleic acid and peanut oil high in oleic acid have been shown to reverse the inhibitory effect upon insulin production and can have a beneficial effect in type 2 diabetes (Vassiliou et al., 2009). Flavonoids have been shown to enhance PPARα expression and normalize blood triglycerides in high-fat-fed diabetic rats (Kaviarasan and Pugalendi, 2009). While trying to develop these targeted products, the developers (food and culinary industries) and the practitioners (mainly registered dietitians and nutritional professional) must consider the regulatory and legal issues associated with nutrigenomics. Since nutrigenomics is a relatively new field, work should be undertaken to develop proper ethical and legal guidelines to protect consumers (Reilly and DeBusk, 2008).
Future Discoveries
Current global trends in food consumption may have an impact on disease progressions observed worldwide. This impact may occur by means of nutrient regulation of genes, or by other unclear mechanisms which are yet to be discovered. There may be opportunities for the food industry to improve public health through modification of processing and production methods, as well as to capitalize on promotion of more positive trends. Nutrigenomic research may uncover the keys to utilizing genetic information to maximize public health and generally benefit the food and culinary industries.
The key to synthesizing this broad spectrum of information into a coherent picture may be the emerging field of nutrigenomics. Through nutrigenomic research, new nutritional regulation of gene expression will hopefully come to light. If specific nutrient regulation of genes closely related to disease onset or progression is identified, new arenas for disease prevention and potential for treatment will come to the foreground of nutritional research and preventive medicine. Treatment and prevention plans based on nutrient consumption will necessarily rely heavily on the available food supply, which, in turn, relies upon the food and culinary industries.
Maximum utilization of the information brought forth by nutrigenomic research is crucial if these industries hope to promote public health.
Jean Getz is a Student, School of Osteopathic Medicine, Michigan State University, East Lansing, MI 48824 (getzjean@msu.edu). Koushik Adhikari, Ph.D., a Member of IFT, is Assistant Professor, Sensory Analysis Center, Dept. of Human Nutrition, Kansas State University, Manhattan, KS 66506 (koushik@ksu.edu). Denis M. Medeiros, Ph.D., R.D., a Professional Member of IFT, is Professor and Head, Dept. of Human Nutrition, Associate Dean for Scholarship and Research, Kansas State University (medeiros@ksu.edu).
References
Afman, L., and Muller, M. 2006. Nutrigenomics: from molecular nutrition to prevention of disease. J. Am. Diet Assoc. 106: 569-576.
Albala, C., Vio, F., Kain, J., and Uauy, R. 2002. Nutrition transition in Chile: determinants and consequences. Public Health Nutr. 5(1A): 123-128.
Allende, A., Tomás-Barberán, F.A., and Gil, M.I. 2006. Minimal processing for healthy traditional foods. Trends Food Sci. Technol. 17: 513-519.
Archibald, A. 2004. Diet Design, One Pair of “Genes” at a Time. Food Product Design. http://www.nutrasolutions.com/Articles/Feature_Article/84ba255543f18010VgnVCM100000f932a8c0. Accessed Sept. 15, 2009.
Aubourg, S., Gallardo, J.M., and Medina, I. 1997. Changes in lipids during different sterilizing conditions in canning albacore (Thunnus alalunga) in oil. Int. J. Food. Sci. Tech. 32: 427-431.
Barsh, G.S., Farooqui, I.S., and O’Rahilly, S. 2000. Genetics of body-weight regulation. Nature. 404: 644-651.
Bouwens, M., Afman, L.A., and Muller, M. 2007. Fasting induces changes in peripheral blood mononuclear cell gene expression profiles related to increases in fatty acid β-oxidation: functional role of peroxisome proliferator-activated protein receptor α in human peripheral blood mononuclear cells. Am. J. Clin. Nutr. 86: 1515-1523.
Bremener, I., and Beattie, J.H. 1990. Metallothionein and the trace minerals. Ann. Rev. Nutr. 10: 63-83.
Clarke, S.D., and Abraham, S. 1992. Gene expression: nutrient control of pre- and posttranscriptional events. FASEB J. 6: 3146-3152.
Croce, C.M. 2008. Oncogenes and cancer: molecular origins of cancer. New Engl. J. Med. 358: 502-511.
DeLuca, L.M. 1991. Retinoids and their receptors in differentiation, embryogenesis, and neoplasia. FASEB J. 5: 2924-2933.
Ducluzeau, P.H., Perretti, N., Laville, M., Andreelli, F., Vega, N., Riou, J.P., and Vidal, H. 2001. Regulation by insulin of gene expression in human skeletal muscle and adipose tissue. Diabetes. 50: 1134-1142.
Ferguson, L.R. 2009. Nutrigenomics approaches to functional foods. J. Amer. Diet. Assoc. 109: 452-458.
Gallegos-Arreola, M.P., García-Ortiz, J.E., Figuera, L.E., Puebla-Pérez, A.M., Morgan-Villela, G., and Zúñiga-González, G.M. 2009. Association of the 677C→T polymorphism in the MTHFR gene with colorectal cancer in Mexican patients. Cancer Genomics Proteomics. 6: 183-188.
German, M.S., Moss, L.G., and Rutter, W.J. 1990. Regulation of insulin gene expression by glucose and calcium in transfected primary islet cultures. J. Biol. Chem. 265: 22063-22066.
Hale, M.B., and Brown, T. 1983. Fatty acids and lipid classes of three underutilized species and changes due to canning. Mar. Fish Rev. 45: 4-6.
Kaviarasan, K., and Pugalendi, K.V. 2009. Influence of flavonoid-rich fraction from Spermacoce hispida seed on PPAR-alpha gene expression, antioxidant redox status, protein metabolism and marker enzymes in high-fat-diet fed STZ diabetic rats. Basic Clin. Physiol. Pharmacol. 20: 141-158.
Komduur, R.H., Korthals, M., and te Molder, H. 2009. The good life: living for health and a life without risks? On a prominent script of nutrigenomics. Br. J. Nutr. 101: 307-316.
Kresten, S., Seydoux, J., Peters, J.M., Gonzalez, F.J., Desvergne, B., and Wahli, W. 1999. Peroxisome proliferator-activated receptor alpha mediates the adaptive response to fasting. J. Clin. Invest. 103: 1489-1498.
Land, H., Parada, L.F., and Weinberg, R.A. 1983. Cellular oncogenes and multistep carcinogenesis. Science. 222: 771-778.
Madrazo, J.A., and Kelly, D.P. 2008. The PPAR trio: regulators of myocardial energy metabolism in health and disease. J. Mol. Cell Cardiol. 44: 968-975.
Martínez de Villarreal, L.E., Delgado-Enciso, I., Valdéz-Leal, R., Ortíz-López, R., Rojas-Martínez, A., Limón-Benavides, C., Sánchez-Peña, M.A., Ancer-Rodríguez, J., Barrera-Saldaña, H.A., and Villarreal-Pérez, J.Z. 2001. Folate levels and N(5),N(10)-methylenetetrahydrofolate reductase genotype (MTHFR) in mothers of offspring with neural tube defects: a case-control study. Arch. Med. Res. 32: 277-282.
Mead, M.N. 2007. Nutrigenomics: the genome-food interface. Environ Health Perspect. 115: A582-A589.
Michaud, S.E., and Renier, G. 2001. Direct regulatory effect of fatty acids on macrophage lipoprotein lipase: potential role of PPARs. Diabetes. 50: 660-666.
Muller, M., and Kersten, S. 2003. Nutrigenomics: goals and strategies. Nat. Rev. Genet. 4: 315-322.
Ordovas, J.M., and Corella, D. 2004. Nutritional Genomics. Annu. Rev. Genomics Hum. Genet. 5: 71–118.
Ortega-Rivas, E. 2007. Processing effects for safety and quality in some predominant food technologies. Cri. Rev. Food Sci. Nutr. 47: 161-173.
Pankow, J.S., Folsom, A.R., Cushman, M., Borecki, I.B., Hopkins, P.N., Eckfeldt, J.H., and Tracy, R.P. 2001. Familial and genetic determinants of systemic markers of inflammation: the NHLBI family heart study. Atherosclerosis. 154: 681-689.
Reilly, P.R., and DeBusk, R.M. 2008. Ethical and legal issues in nutritional genomics. J. Am. Diet. Assoc. 108: 36-40.
Robinson, E., and Grieve, D.J. 2009. Significance of peroxisome proliferatoractivated receptors in the cardiovascular system in health and disease. Pharmacol. Ther. 122: 246-263.
Ronteltap, A., van Trijp, J.C.M., and Renes, R.J. 2009. Consumer acceptance of nutrigenomics-based personalized nutrition. Br. J. Nutr. 101: 132-144.
Samuel, S., and Sitrin, M.D. 2008. Vitamin D’s role in cell proliferation and differentiation. Nutr. Rev. 66(10 Suppl 2): S116-S124.
Schroeder, H.A. 1971. Losses of vitamins and trace minerals resulting from processing and preservation of foods. Am. J. Clin. Nutr. 24: 562-753.
Slavin, J.L., Jacobs, D., and Marquart, L. 2002. Grain processing and nutrition. Crit. Rev. Biotechnol. 21: 49-66.
Vassiliou, E.K., Gonzalez, A., Garcia, C., Tadros, J.H., Chakraborty, G., and Toney, J.H. 2009. Oleic acid and peanut oil high in oleic acid reverse the inhibitory effect of insulin production of the inflammatory cytokine TNF-alpha both in vitro and in vivo systems. Lipids Health Dis. 8: 25-34.
Verreth, W., De Keyzer, D., Pelat, M., Verhamme, P., Ganame, J., Bielicki, J.K., Mertens, A., Quarck, R., Benhabilès, N., Marguerie, G., Mackness, B., Mackness, M., Ninio, E., Herregods, M.C., Balligand, J.L., and Holvoet, P. 2004. Weightloss-associated induction of peroxisome proliferator-activated receptor-alpha and peroxisome proliferator-activated receptor-gamma correlate with reduced atherosclerosis and improved cardiovascular function in obese insulin-resistant mice. Circulation. 110: 3259-3269.
Authors
-
JEAN GETZ, KOUSHIK ADHIKARI, , DENIS MEDEIROS
Categories
-
Food Health Nutrition
-
Food Sciences
-
Food Product Development
-
Functional Foods
-
Food Engineering
-
Food Technology Magazine



